ISMARA: automated modeling of genomic signals as a democracy of regulatory motifs. - Abstract - Europe PMC
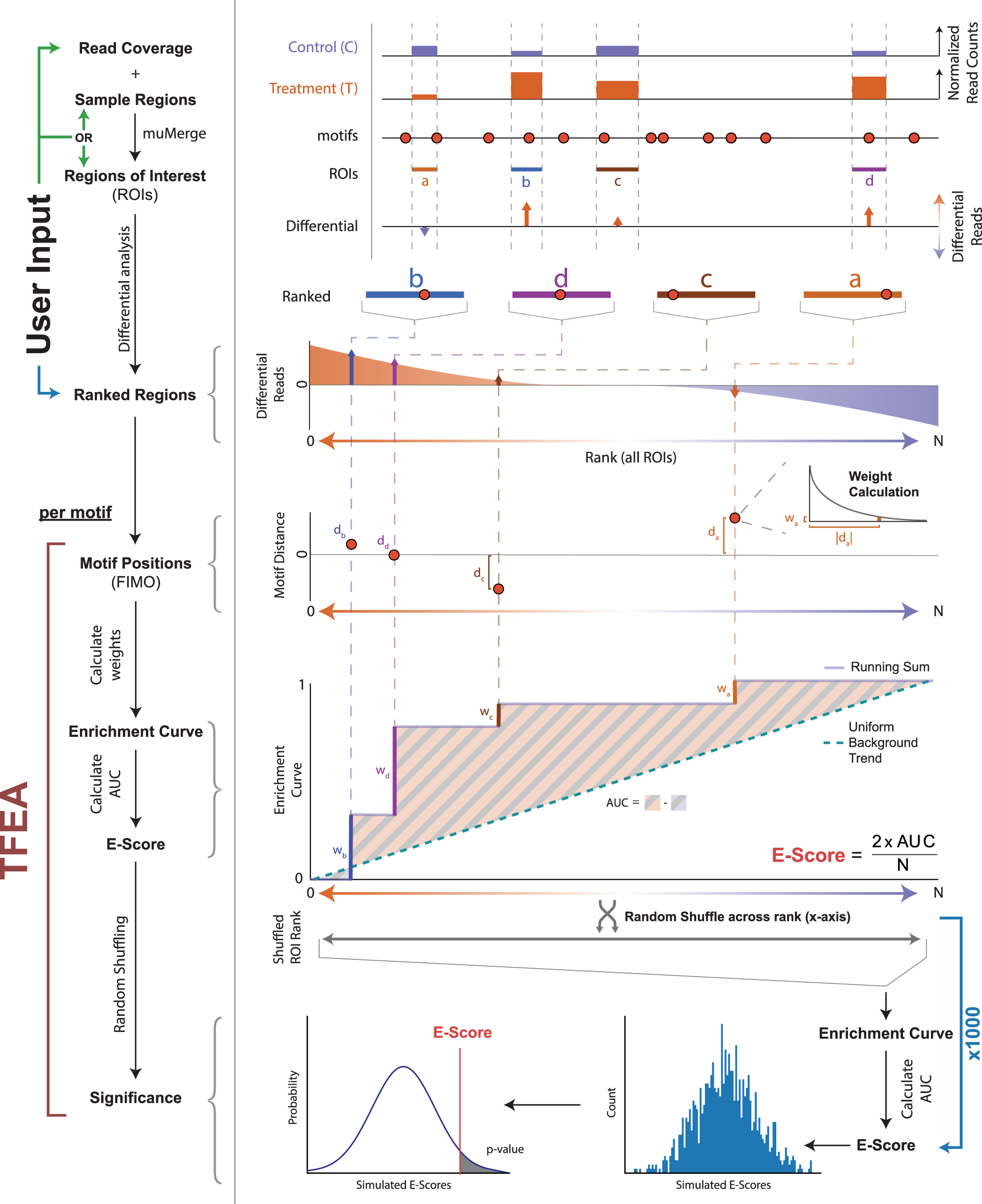
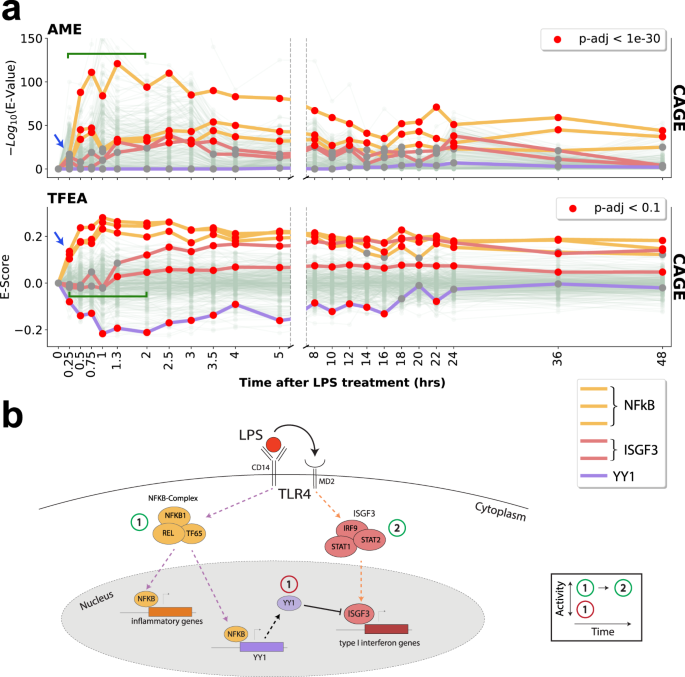
Transcription factor enrichment analysis (TFEA) quantifies the activity of multiple transcription factors from a single experiment | Communications Biology

Bound (Anniversary Edition) (The Nevermore Trilogy) (Volume 2): Mayer, Shannon: 9781540485977: Amazon.com: Books
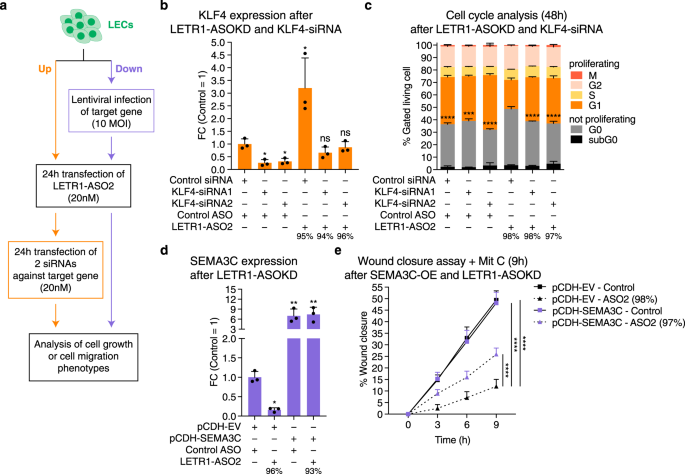
LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C | Nature Communications
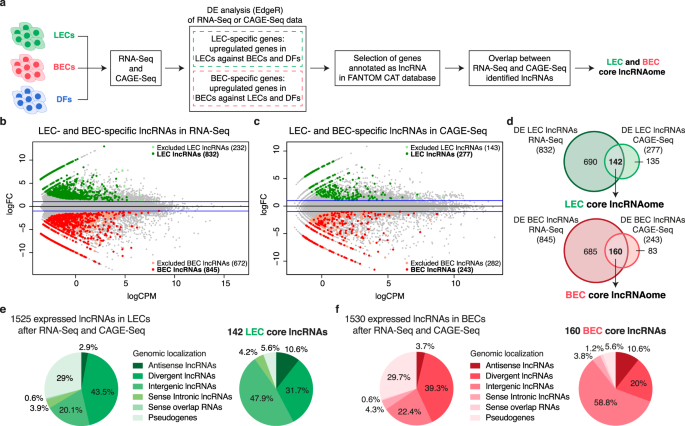
LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C | Nature Communications
GitHub - ismara-unibas/upload-client: Upload client for the ISMARA web-service (www.ismara.unibas.ch)
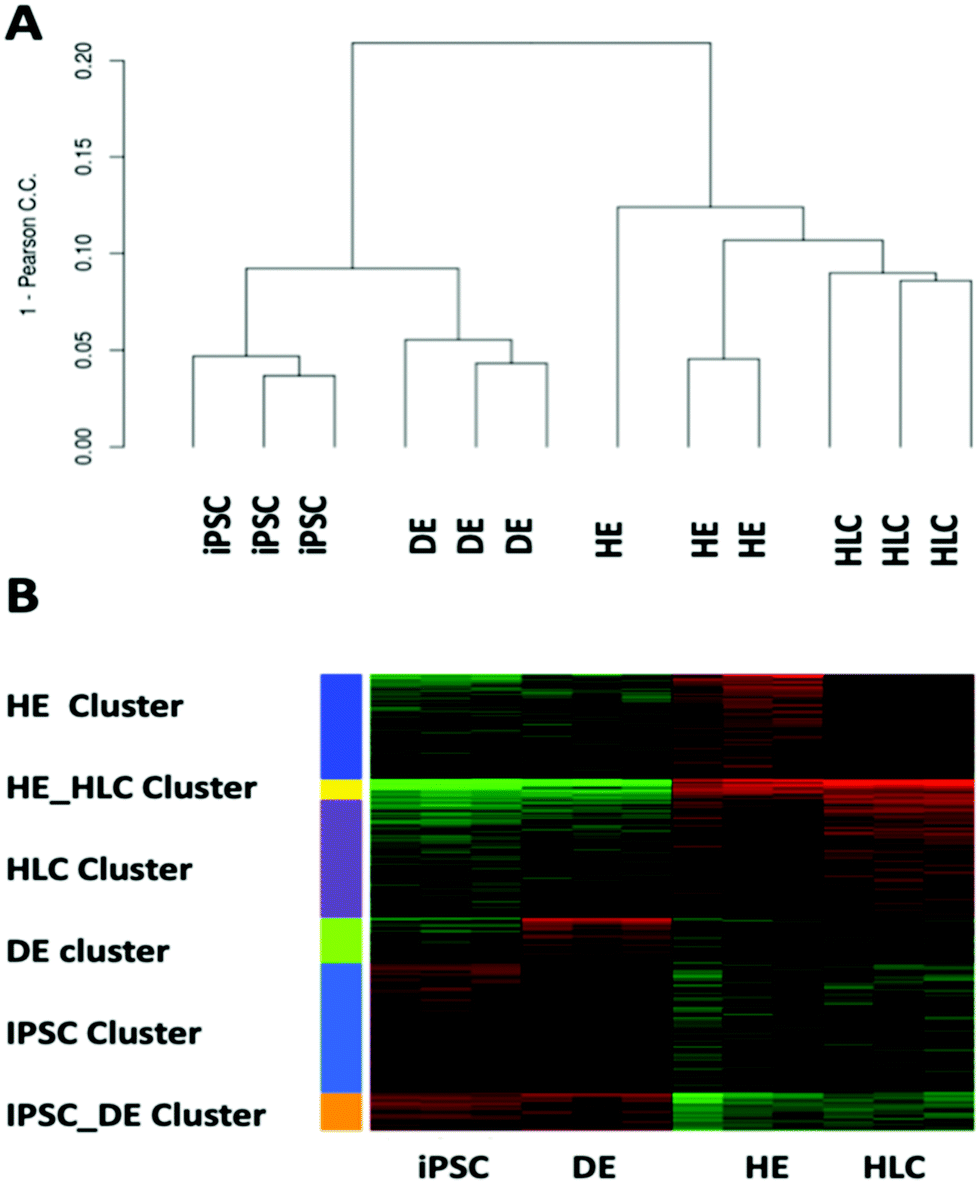
Analysis of the transcription factors and their regulatory roles during a step-by-step differentiation of induced pluripotent stem cells into hepatocy ... - Molecular Omics (RSC Publishing) DOI:10.1039/C9MO00122K
Integrated annotation and analysis of genomic features reveal new types of functional elements and large-scale epigenetic phenom

Deep cap analysis gene expression (CAGE): genome-wide identification of promoters, quantification of their expression, and network inference | BioTechniques

Integrative CAGE and DNA Methylation Profiling Identify Epigenetically Regulated Genes in NSCLC | Molecular Cancer Research | American Association for Cancer Research
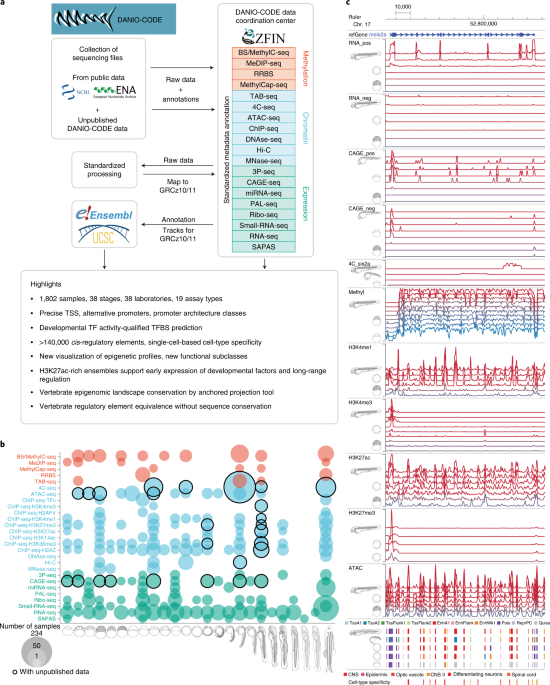
Multiomic atlas with functional stratification and developmental dynamics of zebrafish cis-regulatory elements | Nature Genetics

Integrated annotation and analysis of genomic features reveal new types of functional elements and large-scale epigenetic phenomena in the developing zebrafish | bioRxiv
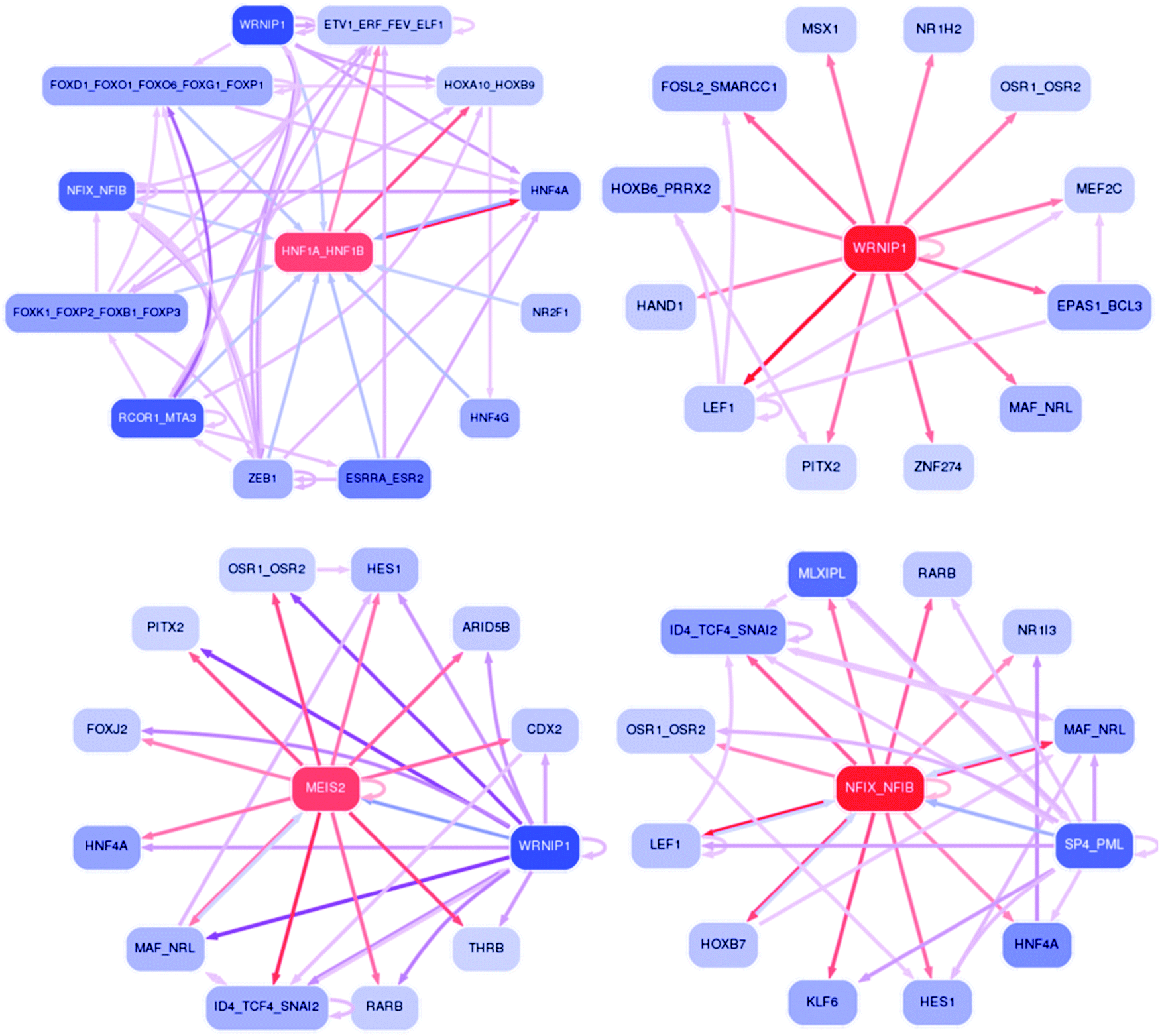
Analysis of the transcription factors and their regulatory roles during a step-by-step differentiation of induced pluripotent stem cells into hepatocy ... - Molecular Omics (RSC Publishing) DOI:10.1039/C9MO00122K